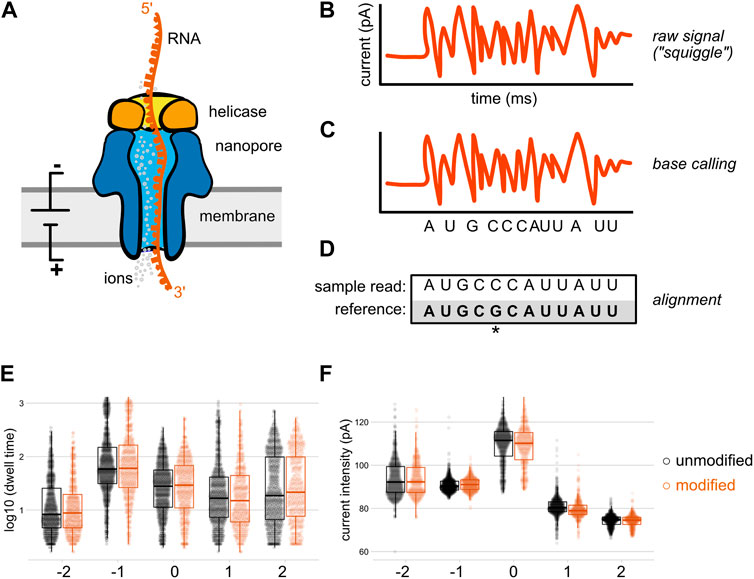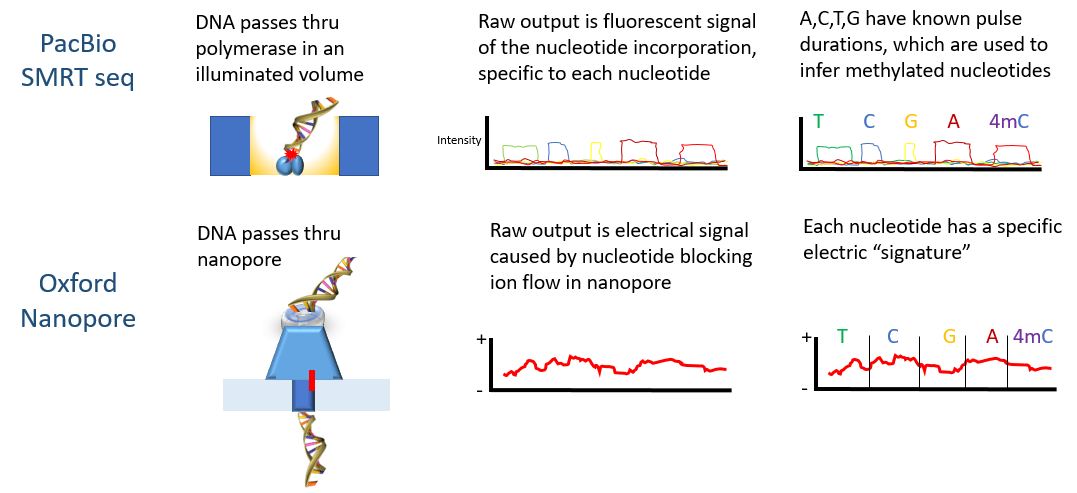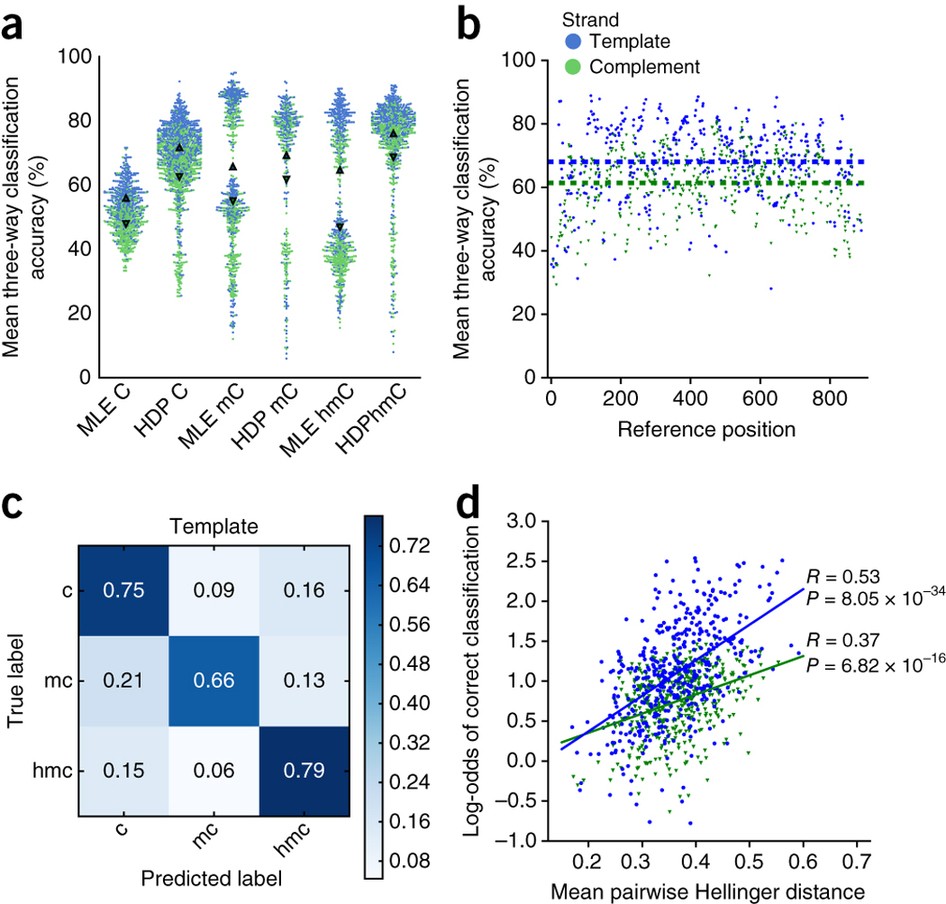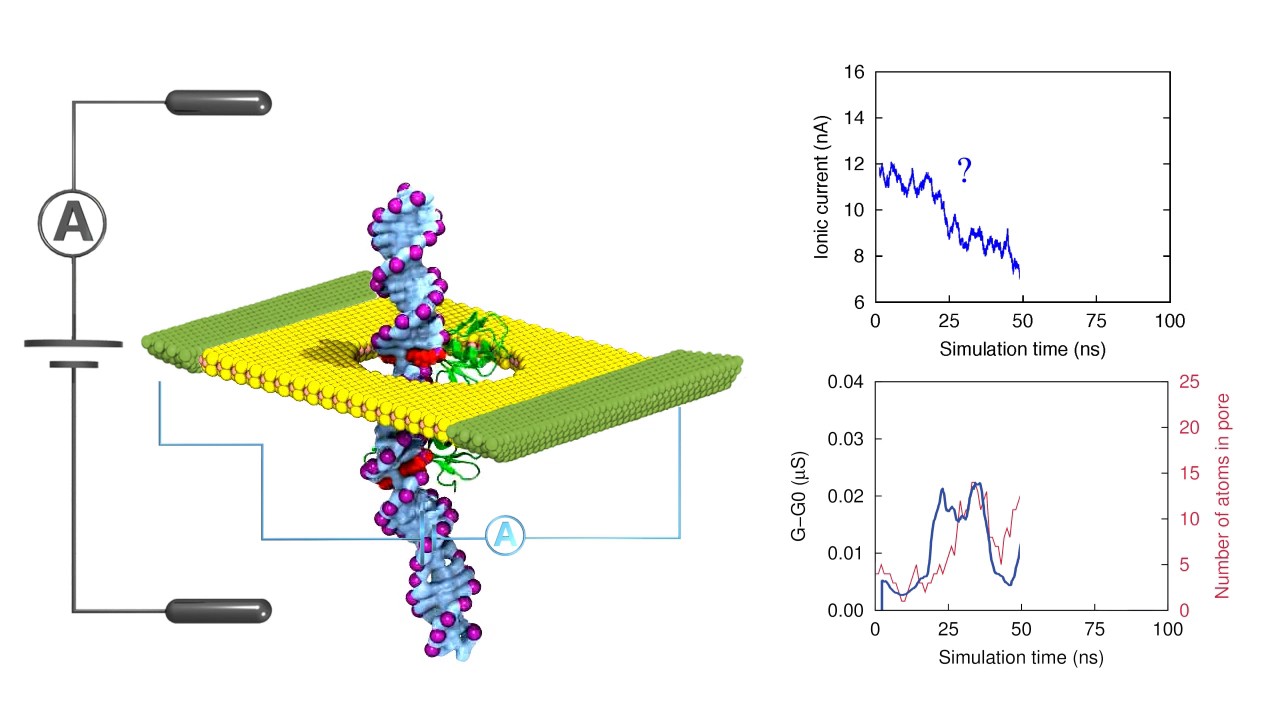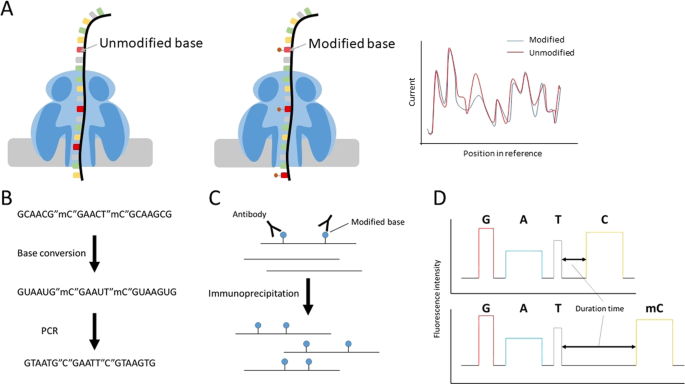
Recent advances in the detection of base modifications using the Nanopore sequencer | Journal of Human Genetics

Discovering multiple types of DNA methylation from bacteria and microbiome using nanopore sequencing | Nature Methods

DNA methylation calling tools for Oxford Nanopore sequencing: a survey and human epigenome-wide evaluation | bioRxiv
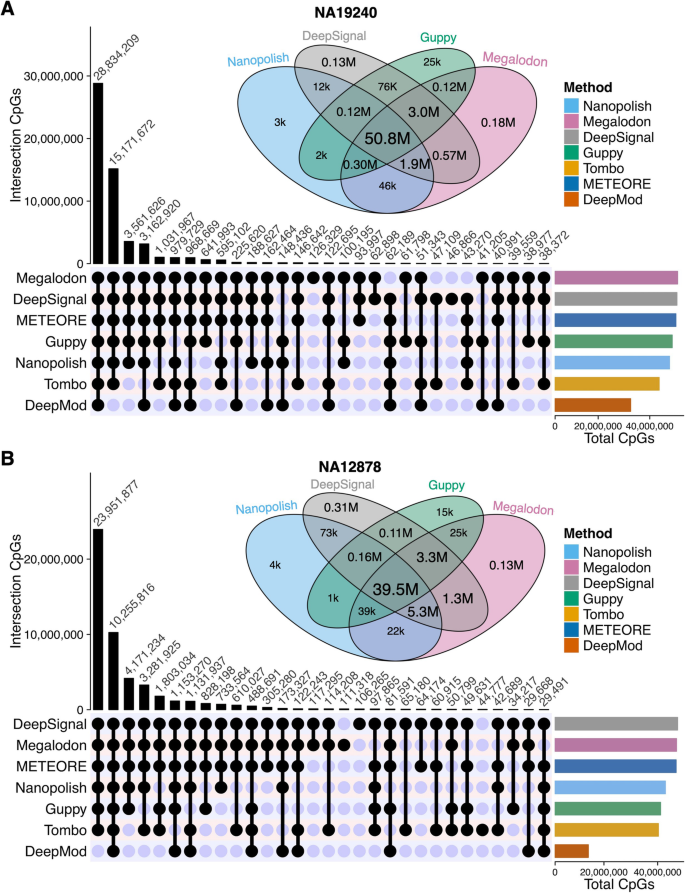
DNA methylation-calling tools for Oxford Nanopore sequencing: a survey and human epigenome-wide evaluation | Genome Biology | Full Text
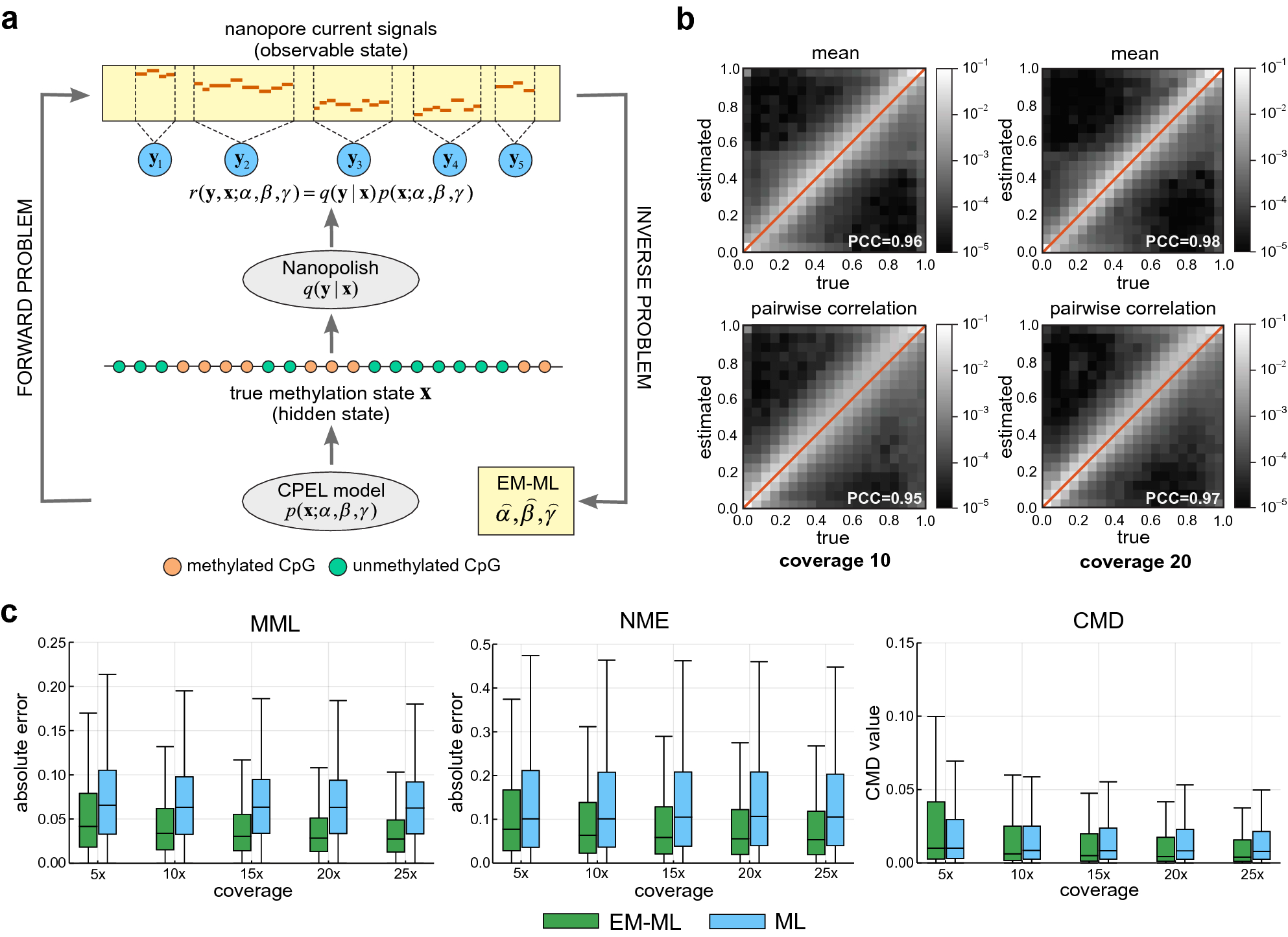
Estimating DNA methylation potential energy landscapes from nanopore sequencing data | Scientific Reports
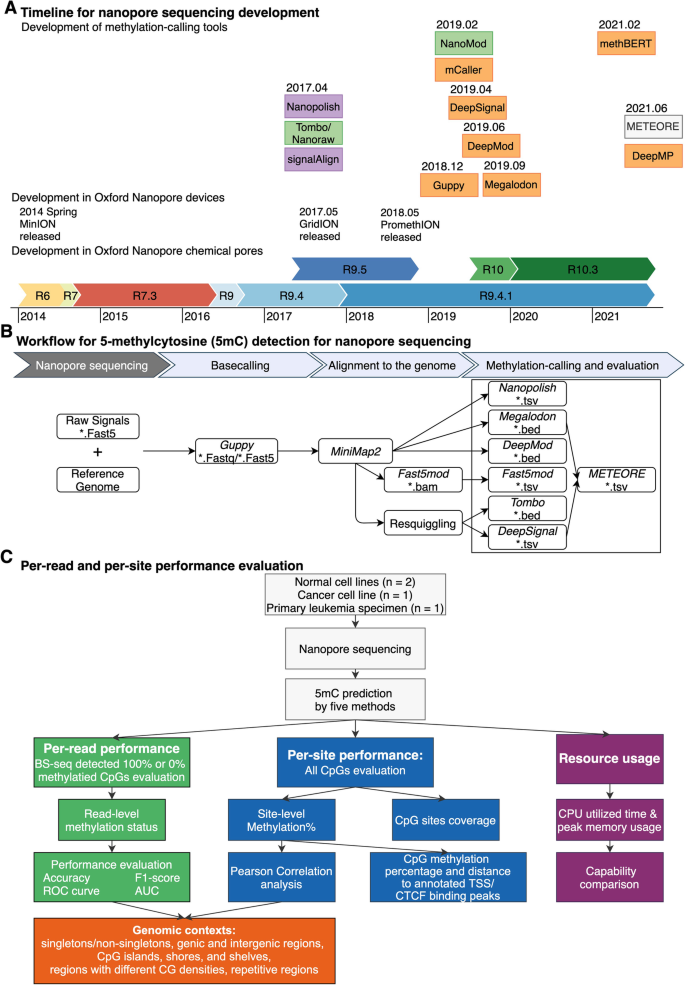
DNA methylation-calling tools for Oxford Nanopore sequencing: a survey and human epigenome-wide evaluation | Genome Biology | Full Text

Discovering and exploiting multiple types of DNA methylation from individual bacteria and microbiome using nanopore sequencing | bioRxiv
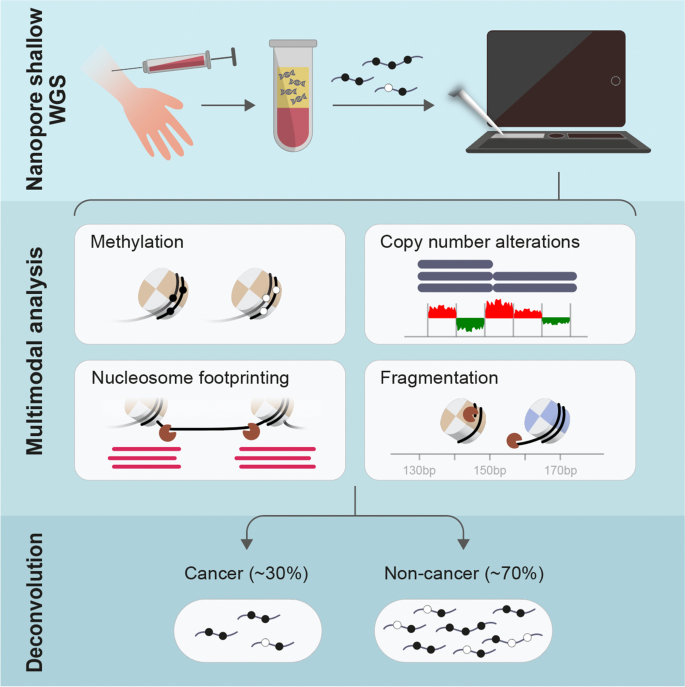
Detecting cell-of-origin and cancer-specific methylation features of cell-free DNA from Nanopore sequencing | Genome Biology | Full Text
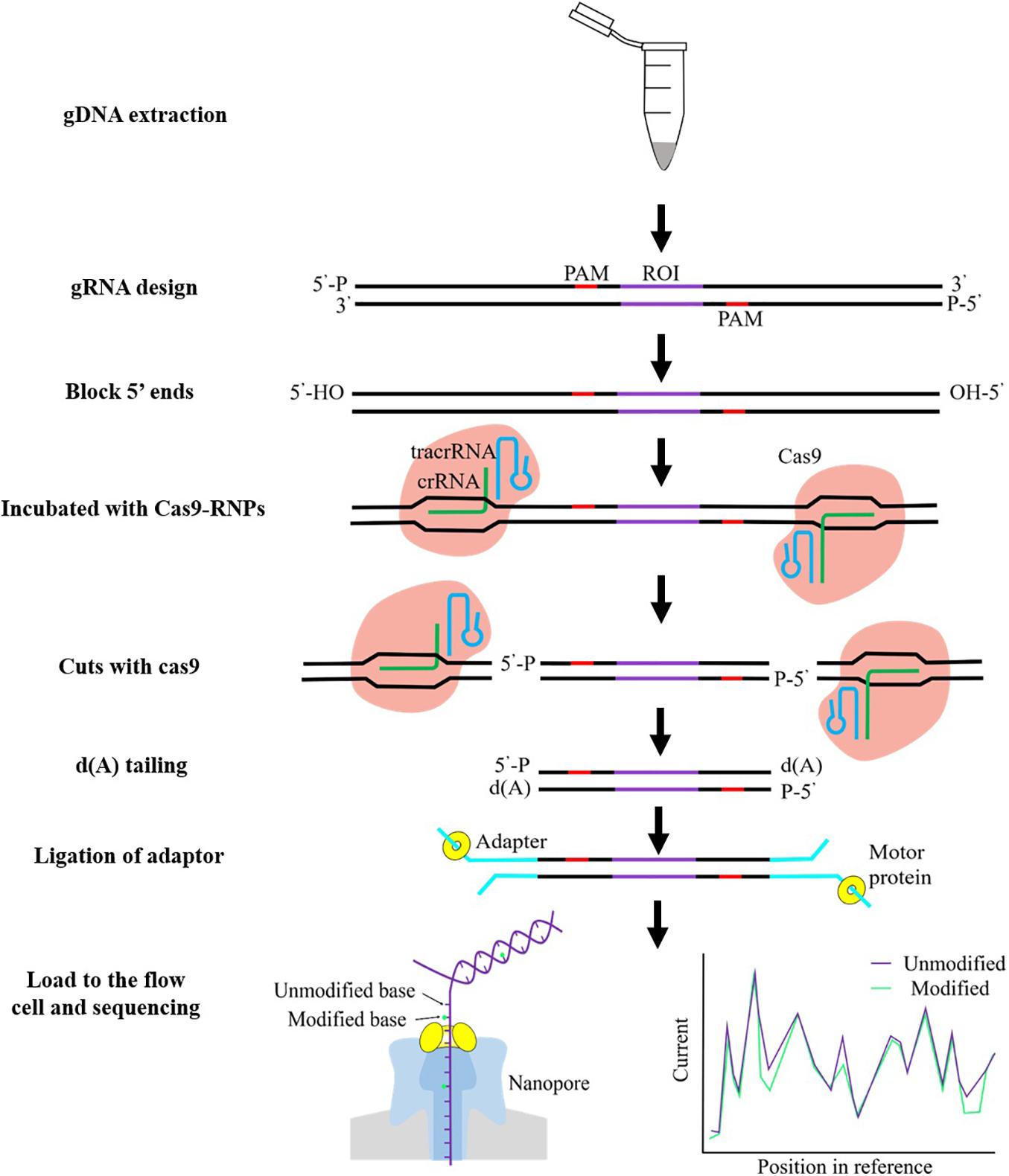
Frontiers | Cancer Biomarkers Discovery of Methylation Modification With Direct High-Throughput Nanopore Sequencing
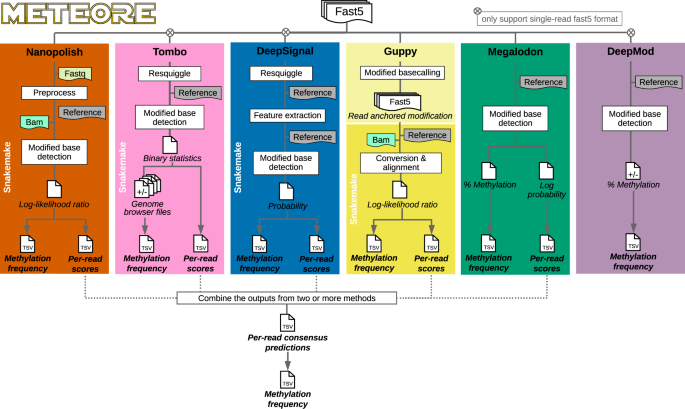
Systematic benchmarking of tools for CpG methylation detection from nanopore sequencing | Nature Communications
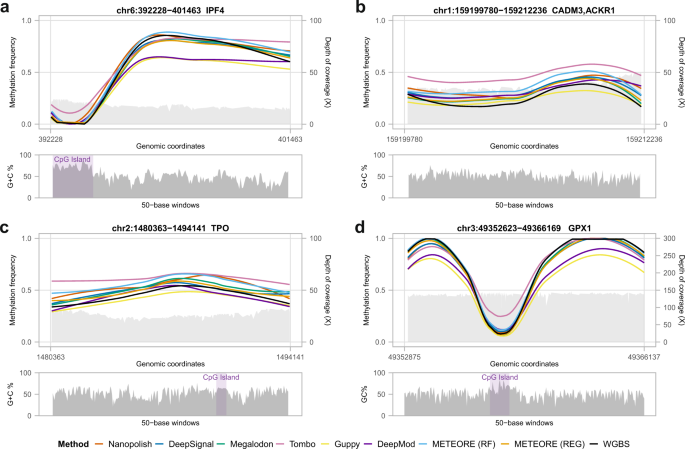
Systematic benchmarking of tools for CpG methylation detection from nanopore sequencing | Nature Communications